Our new paper on ‘measuring integrated information’ is out now, open access, in the journal Entropy. It’s part of a special issue dedicated to integrated information theory.
In consciousness research, ‘integrated information theory’, or IIT, has come to occupy a highly influential and rather controversial position. Acclaimed by some as the most important development in consciousness science so far, critiqued by others as too mathematically abstruse and empirically untestable, IIT is by turns both fascinating and frustrating. Certainly, a key challenge for IIT is to develop measures of ‘integrated information’ that can be usefully applied to actual data. These measures should capture, in empirically interesting and theoretically profound ways, the extent to which ‘a system generates more information than the sum of its parts’. Such measures are also of interest in many domains beyond consciousness, through for example to physics and engineering, where notions of ‘dynamical complexity’ are of more general importance.
Adam Barrett and I have been working towards this challenge for many years, both through approximations of the measure F (‘phi’, central to the various iterations of IIT) and through alternative measures like ‘causal density’. Alongside new work from other groups, there now exist a range of measures of integrated information – yet so far no systematic comparison of how they perform on non-trivial systems.
This is what we provide in our new paper, led by Adam along with Pedro Mediano from Imperial College London.
*
We describe, using a uniform notation, six different candidate measures of integrated information (among which we count the related measure of ‘causal density’). We set out the intuitions behind each, and compare their properties across a series of criteria. We then explore how they behave on a variety of network models, some very simple, others a little bit more complex.
The most striking finding is that the measures all behave very differently – no two measures show consistent agreement across all our analyses. Here’s an example:
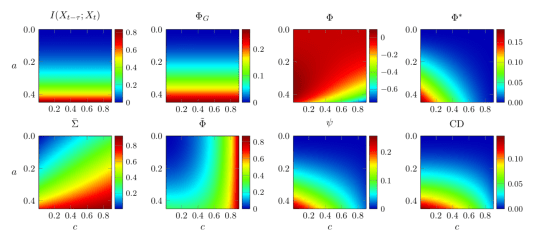
Diverse behavior of measures of integrated information. The six measures (plus two control measures) are shown in terms of their behavior on a simple 2-node network animated by autoregressive dynamics.
At first glance this seems worrying for IIT since, ideally, one would want conceptually similar measures to behave in similar ways when applied to empirical test-cases. Indeed, it is worrying if existing measures are used uncritically. However, by rigorously comparing these measures we are able to identify those which better reflect the underlying intuitions of ‘integrated information’, which we believe will be of some help as these measures continue to be developed and refined.
Integrated information, along with related notions of dynamical complexity and emergence, are likely to be important pillars of our emerging understanding of complex dynamics in all sorts of situations – in consciousness research, in neuroscience more generally, and beyond biology altogether. Our new paper provides a firm foundation for the future development of this critical line of research.
*
One important caveat is necessary. We focus on measures that are, by construction, applicable to the empirical, or spontaneous, statistically stationary distribution of a system’s dynamics. This means we depart, by necessity, from the supposedly more fundamental measures of integrated information that feature in the most recent iterations of IIT. These recent versions of the theory appeal to the so-called ‘maximum entropy’ distribution since they are more interested in characterizing the ‘cause-effect structure’ of a system than in saying things about its dynamics. This means we should be very cautious about taking our results to apply to current versions of IIT. But, in recognizing this, we also return to where we started in this post. A major issue for the more recent (and supposedly more fundamental) versions of IIT is that they are extremely challenging to operationalize and therefore to put to an empirical test. Our work on integrated information departs from ‘fundamental’ IIT precisely because we prioritise empirical applicability. This, we think, is a feature, not a bug.
*
All credit for this study to Pedro Mediano and Adam Barrett, who did all the work. As always, I’d like to thank the Dr. Mortimer and Theresa Sackler Foundation, and the Canadian Institute for Advanced Research, Azrieli Programme in Brain, Mind, and Consciousness, for their support. The paper was published in Entropy on Christmas Day, which may explain why some of you might’ve missed it! But it did make the cover, which is nice.
*
Mediano, P.A.M., Seth, A.K., and Barrett, A.B. (2019). Measuring integrated information. Comparison of candidate measures in theory and in simulation. Entropy, 21:17